Acessing model data using intake-esm#
To access the CMIP6 model data and CESM-PPE data we created intake catalogs to help browse and subset the data. The catalogs are stored in shared folder /mnt/craas2-ns9988k/data/catalogs/
# Importing the required packages
import intake
import xarray as xr
import intake_esm
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
Reading and browsing the catalog#
# Open the catalog
# col = intake.open_esm_datastore('https://storage.googleapis.com/cmip6/pangeo-cmip6.json') # Remote Pangeo
# they are also catalogs abailable in the shared read-only data-space.
col = intake.open_esm_datastore('/mnt/craas2-ns9988k/data/catalogs/cmip6.json') # Local data (+pangeo) stored on NIRD
col
cmip6 catalog with 155 dataset(s) from 536945 asset(s):
| unique | |
|---|---|
| variable_id | 583 |
| table_id | 24 |
| source_id | 75 |
| experiment_id | 94 |
| member_id | 190 |
| grid_label | 11 |
| time_range | 9100 |
| activity_id | 18 |
| institution_id | 35 |
| version | 577 |
| path | 536945 |
| dcpp_init_year | 0 |
| derived_variable_id | 0 |
Under the hood intake-esm uses a large table csv, which contains some metadata and the paths of where to find it. The data can be stored both locally or in a remote cloud storage i.e pangeo-cloud.
Note
Since the paths listed in the csv table are absolute, notebook starting from the same catalog can be run from any directory, without needing to change the in side the notebook paths.
Browsing the catalog: Comparing the change in AOD over the historical period across CMIP6 models#
The keywords when browsing is columns of the table, e.g. variable_id, source_uid etc. To list all the keys available for a given key word you can use the col.unique()['<keyword>'] function.
col = col.search(
variable_id='od550aer',
experiment_id='histSST'
)
col
cmip6 catalog with 9 dataset(s) from 190 asset(s):
| unique | |
|---|---|
| variable_id | 1 |
| table_id | 1 |
| source_id | 9 |
| experiment_id | 1 |
| member_id | 4 |
| grid_label | 3 |
| time_range | 185 |
| activity_id | 2 |
| institution_id | 9 |
| version | 9 |
| path | 190 |
| dcpp_init_year | 0 |
| derived_variable_id | 0 |
Models available for this request:
col.unique()['source_id']
['CNRM-ESM2-1',
'MPI-ESM-1-2-HAM',
'MIROC6',
'UKESM1-0-LL',
'MRI-ESM2-0',
'GISS-E2-1-G',
'CESM2-WACCM',
'GFDL-ESM4',
'EC-Earth3-AerChem']
Loading the data and plotting#
Warning
The catalog can be huge, always query and subset catalog before loading the data. Preferably down to level of experiment and variable.
The .to_dataset_dict function can accept an optional preprocessing function which can be used to harmonize the datasets, temporal resampling, or slicing.
Below we made a preprocessing function for resampling the data into annual means and calculate the global average to allow for easily plotting the time series.
def resample_calculate_and_global_avg(ds):
"""
Function to resample the data to annual mean and calculate the global average.
"""
ds=ds.resample(time='YE').mean() # Resample to annual mean
ds = ds.drop_vars(['lat_bnds','lon_bnds'], errors='ignore') # create conflict when calculating global average
weights = np.cos(np.deg2rad(ds.lat)) # Make weighted global average
ds_weighted = ds.weighted(weights)
weighted_mean = ds_weighted.mean(dim=['lon','lat'])
return weighted_mean
The processing can be done in parallel using dask, which can speed things up a bit.
The dask cluster below is constrained within the resource each user have available.
The corresponding dask cluster panel can be viewed by clicking on the dask icon on the left-side panel.
from dask.distributed import Client, LocalCluster
client = Client(LocalCluster(n_workers=4, threads_per_worker=1, memory_limit='12GB'))
Don’t forget to close the client after you done with client.shutdown().
# dataset_dict = col.to_dataset_dict()
dataset_dict = col.to_dataset_dict(xarray_open_kwargs={'use_cftime':True, 'chunks':{'time':48}},
aggregate=True,
preprocess=resample_calculate_and_global_avg,
skip_on_error=True,
xarray_combine_by_coords={'combine_attrs': 'override'}
)
--> The keys in the returned dictionary of datasets are constructed as follows:
'activity_id.institution_id'
Without the providing a preprocess function the time aggregated output as dictionary with grouped by the following keys "activity_id", "institution_id", "source_id", "experiment_id", "table_id", "grid_label".
col.to_dataset_dict(xarray_open_kwargs={'use_cftime':True, 'chunks':{'time':48}},
aggregate=True,
skip_on_error=True,
xarray_combine_by_coords={'combine_attrs': 'override'}
)
Preprocess dictionary only contain the time series, which is easy to loop over and plot for each model.
dataset_dict
{'AerChemMIP.NASA-GISS': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p3f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: GISS-E2-1-G
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p3f1
intake_esm_attrs:grid_label: gn
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: NASA-GISS
intake_esm_attrs:version: v20191120
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.NASA-GISS,
'AerChemMIP.CNRM-CERFACS': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p1f2'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes: (12/14)
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: CNRM-ESM2-1
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f2
... ...
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: CNRM-CERFACS
intake_esm_attrs:version: v20190621
intake_esm_attrs:path: /mnt/craas2-ns9988k-dl-ns9560k/ESGF/CMI...
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.CNRM-CERFACS,
'AerChemMIP.NCAR': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i2p1f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: CESM2-WACCM
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i2p1f1
intake_esm_attrs:grid_label: gn
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: NCAR
intake_esm_attrs:version: v20190531
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.NCAR,
'AerChemMIP.NOAA-GFDL': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p1f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: GFDL-ESM4
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f1
intake_esm_attrs:grid_label: gr1
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: NOAA-GFDL
intake_esm_attrs:version: v20180701
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.NOAA-GFDL,
'AerChemMIP.MRI': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
wavelength float64 8B 550.0
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p1f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes: (12/14)
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: MRI-ESM2-0
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f1
... ...
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: MRI
intake_esm_attrs:version: v20200207
intake_esm_attrs:path: /mnt/craas2-ns9988k-dl-ns9560k/ESGF/CMI...
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.MRI,
'AerChemMIP.HAMMOZ-Consortium': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
wavelength float64 8B 550.0
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p1f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: MPI-ESM-1-2-HAM
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f1
intake_esm_attrs:grid_label: gn
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: HAMMOZ-Consortium
intake_esm_attrs:version: v20190628
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.HAMMOZ-Consortium,
'AerChemMIP.MOHC': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
wavelength float64 8B 550.0
* time (time) object 1kB 1850-12-30 00:00:00 ... 2014-12-30 00:00:00
* member_id (member_id) object 8B 'r1i1p1f2'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: UKESM1-0-LL
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f2
intake_esm_attrs:grid_label: gn
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: MOHC
intake_esm_attrs:version: v20190902
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.MOHC,
'AerChemMIP.MIROC': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
wavelength float64 8B 550.0
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p1f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 4), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: MIROC6
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f1
intake_esm_attrs:grid_label: gn
intake_esm_attrs:activity_id: AerChemMIP
intake_esm_attrs:institution_id: MIROC
intake_esm_attrs:version: v20190828
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: AerChemMIP.MIROC,
'CMIP.EC-Earth-Consortium': <xarray.Dataset> Size: 3kB
Dimensions: (time: 165, member_id: 1)
Coordinates:
wavelength float64 8B 550.0
* time (time) object 1kB 1850-12-31 00:00:00 ... 2014-12-31 00:00:00
* member_id (member_id) object 8B 'r1i1p1f1'
Data variables:
od550aer (member_id, time) float64 1kB dask.array<chunksize=(1, 1), meta=np.ndarray>
Attributes:
intake_esm_vars: ['od550aer']
intake_esm_attrs:variable_id: od550aer
intake_esm_attrs:table_id: AERmon
intake_esm_attrs:source_id: EC-Earth3-AerChem
intake_esm_attrs:experiment_id: histSST
intake_esm_attrs:member_id: r1i1p1f1
intake_esm_attrs:grid_label: gn
intake_esm_attrs:activity_id: CMIP
intake_esm_attrs:institution_id: EC-Earth-Consortium
intake_esm_attrs:version: v20210309
intake_esm_attrs:_data_format_: netcdf
intake_esm_dataset_key: CMIP.EC-Earth-Consortium}
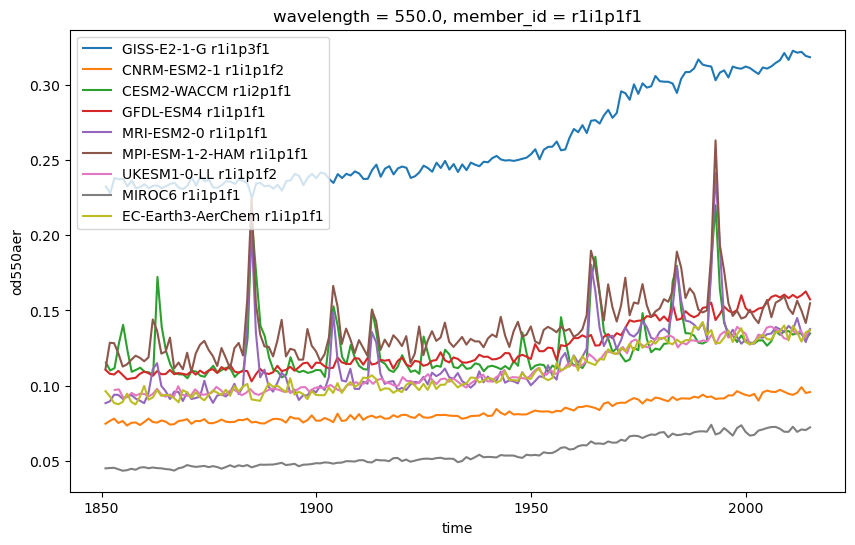
fig,ax = plt.subplots(figsize=(10,6))
for datakey, data in dataset_dict.items(): # Looping over ever varable in the dictionary
source_id = data.attrs['intake_esm_attrs:source_id']
member_id = data.attrs['intake_esm_attrs:member_id']
data['od550aer'].isel(member_id=0).plot(label=f'{source_id} {member_id}', ax=ax)
ax.legend()
<matplotlib.legend.Legend at 0x7fb75ca60410>
Exporting a subset of the catalog#
Most likely you will only be analyzing a small subset of experiments the model experiment and it could be beneficial to work with a reduced catalog.
Below we will make a subset that only contain the information related to histSST and histSST-piaer, and only the absorption optical depth (od550abs) and total optical depth (od550aer).
import os
col = intake.open_esm_datastore('./cmip6.json') # Local data stored on NIRD catalog file in this folder
col = col.search(
experiment_id=['histSST', 'histSST-piaer'],
variable_id = ['od550abs', 'od550aer']
)
col
cmip6 catalog with 9 dataset(s) from 190 asset(s):
| unique | |
|---|---|
| variable_id | 1 |
| table_id | 1 |
| source_id | 9 |
| experiment_id | 1 |
| member_id | 4 |
| grid_label | 3 |
| time_range | 185 |
| activity_id | 2 |
| institution_id | 9 |
| version | 9 |
| path | 190 |
| dcpp_init_year | 0 |
| derived_variable_id | 0 |
Then when we are happy with the selection the catalog can be exported as follows:
col.serialize(name='histSST-AerChemMIP',catalog_type='file',directory='~/')
Successfully wrote ESM catalog json file to: file:///home/fc-3auid-3a41f39e02-2da958-2d4d3b-2d84bd-2d4a2f2886a11b//histSST-AerChemMIP.json
col = intake.open_esm_datastore('~/histSST-AerChemMIP.json')
col
histSST-AerChemMIP catalog with 9 dataset(s) from 190 asset(s):
| unique | |
|---|---|
| variable_id | 1 |
| table_id | 1 |
| source_id | 9 |
| experiment_id | 1 |
| member_id | 4 |
| grid_label | 3 |
| time_range | 185 |
| activity_id | 2 |
| institution_id | 9 |
| version | 9 |
| path | 190 |
| dcpp_init_year | 0 |
| derived_variable_id | 0 |
